Understanding epistasis in linkage analysis : the Kap and lks2 loci in the Oregon Wolfe Barley poputlation | Semantic Scholar
Molecular mapping of quantitative trait loci (QTL) controlling aluminium tolerance in wheat and barley”

PDF) High genetic diversity revealed in barley ( Hordeum vulgare ) collected from small-scale farmer ' s fields in | Jihad Orabi - Academia.edu

Banding pattern of the nSSR loci HVM54 and HVM03 for two barley (B1 and... | Download Scientific Diagram

Banding pattern of the nSSR loci HVM54 and HVM03 for two barley (B1 and... | Download Scientific Diagram

Characterization of the ten transferable microsatellite markers across... | Download Scientific Diagram
MICROSATELLITE ANALYSIS OF GENETIC DIVERSITY OF WILD BARLEY (HORDEUM VULGARE SUBSP. SPONTANEUM) USING DIFFERENT SAMPLING METHODS

PDF) Selection footprints in barley breeding lines detected by combining genotyping-by-sequencing with reference genome information | Elsayed Mansour - Academia.edu

PDF) Development of gene-specific markers for acid soil/aluminium tolerance in barley (Hordeum vulgare L.) | Meixue Zhou - Academia.edu
Determination of QTLs Associated with Agronomic and Physiological Traits under Normal and Salinity Conditions in Barley

Development of gene-specific markers for acid soil/aluminium tolerance in barley (Hordeum vulgare L.) | SpringerLink

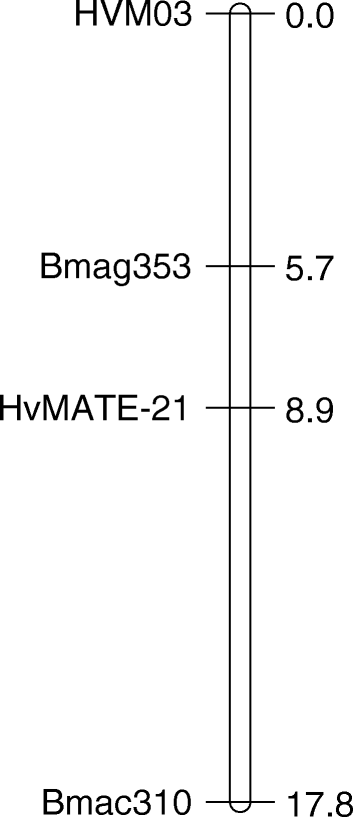
Linkage map of the gene-specific marker Cit7 on chromosome 4H in the... | Download Scientific Diagram
Selection footprints in barley breeding lines detected by combining genotyping-by-sequencing with reference genome information

Cross transferability of barley nuclear SSRs to pearl millet genome provides new molecular tools for genetic analyses and marker assisted selection
Analysis of the efficiency in the Spanish National Barley Breeding Program. Past results and prospects for future improvements u